This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Phylogeny
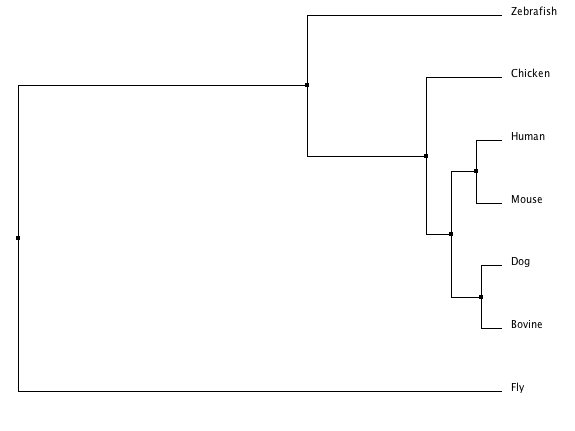
The following trees were created using T-COFFEE (1,2), Phylogeny.fr (6,7,8,9,10,11,12), MUSCLE (5,6), and ClustalW2 (3,4). Each tree is made from an alignment of the Fgfr2 protein in humans to its homologous proteins in mouse (Mus musculus), fly (Drosphilia melanogaster), chicken (Gallus gallus), dog (Canis lupus), bovine (Bos taurus), and zebrafish (Danio rerio).
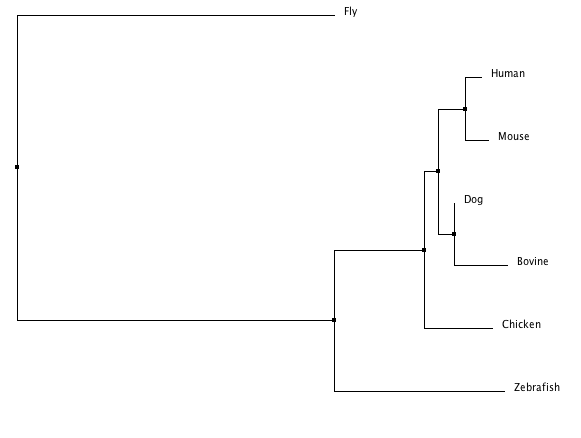
T-COFFEE Phylogeny
These trees include a neighbor joining and an average distance tree. All default settings were used.
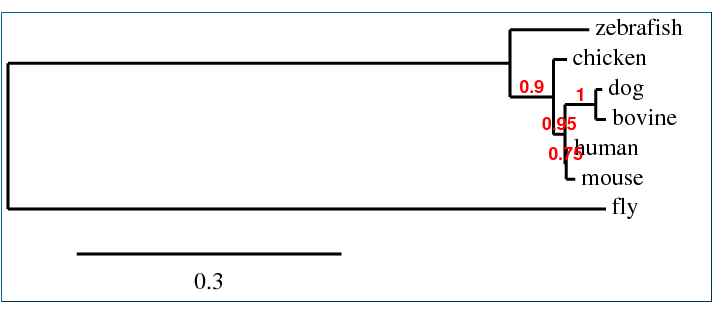
Phylogeny.fr Phylogeny
The following tree was produced on "One Click Mode". All other default settings were used
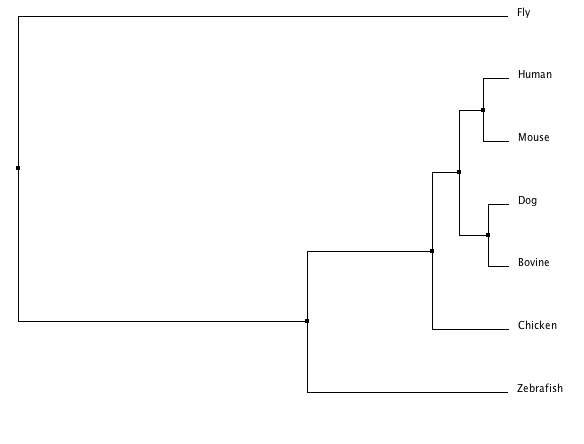
MUSCLE trees
These trees include a neighbor joining and an average distance tree. All default settings were used.
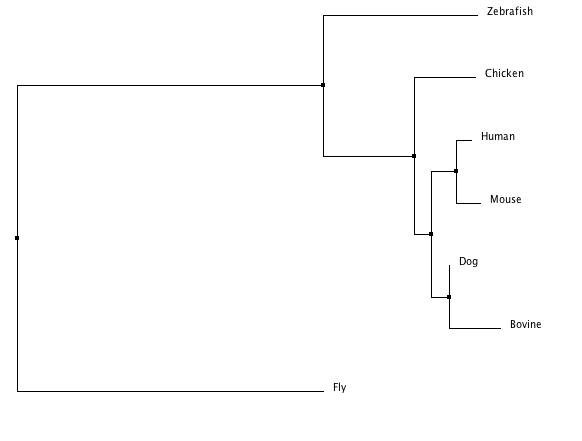
ClustalW tree
The following tree was produced using all default settings.

Analysis
The ClustalW tree suggests the human and mouse protein homologs are very close as well as those of the bovine and dog. It includes a clade with zebrafish, fly, and chicken suggesting those three are more closely related to one another than to the other four species listed. The other trees suggests the Fgfr2 protein in the fly is an outlier among these homologs. This idea is consistent with the T-COFFEE alignment scores (see Homology) which indicated the fly, Drosophilia melanogaster was least similar to the other species in terms of this protein. The Phylogeny.fr tree also provides bootstrap numbers which indicate how likely the phylogenic tree is. There we see the nodes are all .75 or higher of a possible 1 so that tree (and the MUSCLE and T-COFFEE trees which are identical) is a likely analysis of the evolution of this protein.
References
1. Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 2011 Jul;39(Web Server issue):W13-7. Epub 2011 May 9
PMID: 21558174
2.Notredame C, Higgins DG, Heringa J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J Mol Biol. 2000 Sep 8;302(1):205-17
PMID: 10964570
3. Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG. (2007). Clustal W and Clustal X version 2.0. Bioinformatics, 23, 2947-2948
PMID: 17846036
4. Thompson JD, Higgins DG, Gibson TJ. (1994). CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res., 22, 4673-4680
PMID: 7984417
5. Edgar, RC (2004). MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5 (1): 113.
PMID: 15318951
6. Edgar, RC (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Research 32 (5): 1292-97.
PMID: 15034147
7. Dereeper A., Audic S., Claverie J., Blanc G. (2010). BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. Jan 12;10:8 PMID:20067610
8. Dereeper A.*, Guignon V.*, Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.F., Guindon S., Lefort V., Lescot M., Claverie J.M., Gascuel O. (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008 Jul 1;36(Web Server issue):W465-9
PMID:18424797
9. Castresana J. (2004). Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 17(4):540-52
PMID:10742046
10. Guindon S., Gascuel O. (2003). A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 52(5):696-704
PMID:14530136
11. Anisimova M., Gascuel O. (2006). Approximate likelihood ratio test for branchs: A fast, accurate and powerful alternative. Syst Biol. 55(4):539-52.
PMID:16785212
12. Chevenet F., Brun C., Banuls AL., Jacq B., Chisten R. (2006). TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinformatics. 7:439.
PMID:17032440
PMID: 21558174
2.Notredame C, Higgins DG, Heringa J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J Mol Biol. 2000 Sep 8;302(1):205-17
PMID: 10964570
3. Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG. (2007). Clustal W and Clustal X version 2.0. Bioinformatics, 23, 2947-2948
PMID: 17846036
4. Thompson JD, Higgins DG, Gibson TJ. (1994). CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res., 22, 4673-4680
PMID: 7984417
5. Edgar, RC (2004). MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5 (1): 113.
PMID: 15318951
6. Edgar, RC (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Research 32 (5): 1292-97.
PMID: 15034147
7. Dereeper A., Audic S., Claverie J., Blanc G. (2010). BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. Jan 12;10:8 PMID:20067610
8. Dereeper A.*, Guignon V.*, Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.F., Guindon S., Lefort V., Lescot M., Claverie J.M., Gascuel O. (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008 Jul 1;36(Web Server issue):W465-9
PMID:18424797
9. Castresana J. (2004). Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 17(4):540-52
PMID:10742046
10. Guindon S., Gascuel O. (2003). A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 52(5):696-704
PMID:14530136
11. Anisimova M., Gascuel O. (2006). Approximate likelihood ratio test for branchs: A fast, accurate and powerful alternative. Syst Biol. 55(4):539-52.
PMID:16785212
12. Chevenet F., Brun C., Banuls AL., Jacq B., Chisten R. (2006). TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinformatics. 7:439.
PMID:17032440